Xenopus tropicalis genome data
Experimental annotation and analysis of epigenetic regulation
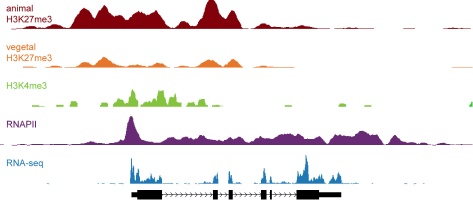
Epigenetic mechanisms set apart the active and inactive regions in the genome of multicellular organisms to produce distinct cell fates during embryogenesis. Transcription profiles, chromatin state maps and genome-wide location analysis of regulatory factors can be used to analyze early development and its early cell lineage decisions. The profiles provide quantitative data on epigenetic regulation and transcriptional activity in the early vertebrate embryo. Moreover, the data can be used to identify promoters, enhancers and transcribed regions and to verify or improve gene annotation.
Xenopus track hub
We have made a track data hub for the UCSC genome browser. This enables easy access to all data without the need to download large files. Please visit our track hub page for more information.
Downloads
Visualization and annotation tracks can be used as custom tracks in the UCSC Genome Browser using the links below. The visualization tracks are available in wiggle format and represent summed read counts in 10 bp windows (ChIP-seq, RNA-seq) or at single base pair resolution (TSS-seq). The XTEV (X. tropicalis experimentally validated gene models) and CpG island annotation tracks are available in BED format.
| File name legend | |
| CpGI | CpG island |
| DNAme | DNA methylation as assayed by methylCap-sequencing (salt fractions, mM) |
| H3K4me3 | Histone H3 lysine 4 trimethylation (gene 5' ends) |
| H3K27me3 | Histone H3 lysine 27 trimethylation (Polycomb repression) |
| RNA-seq | Sequencing of polyA+ RNA (cDNA) |
| RNAPII | RNA polymerase II (associated with DNA) |
| TBP | TATA binding protein (at promoters) |
| TSS-seq | Transcription start site sequencing (based on 5' cap structure of RNA) |
| Xtev | Xenopus tropicalis experimentally validated gene models |
| mostconserved | UCSC phastCons "most conserved" track, liftover from JGI4.1 to 7.1 using GMAP |
| animal, vegetal | animal and vegetal hemispheres of dissected embryos (st.12) |
| "tracks" | bigWig file referencing multiple wiggle tracks |
Genome assembly Xentr v7.1
The "tracks" file below is a bigWig file that can be used to visualize H3K4me3, TBP, H3K4me1, RNAPII, H3K27me3, RNAseq, MethylCap and Input tracks of multiple stages (stages 9, 12, 16 and 30) at the genome browser at NIMR. Alternatively, browse genes for H3K4me3 and H3K27me3 profiles at our gene profile page.
|
|||||||||||||||||
|
|
|||||||||||||||||
Genome assembly Joint Genome Institute Xentr v4.1 (xenTro2, August 2005).
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
Except for unpublished tracks (UNPUB), the raw data have been deposited in NCBI's Gene Expression Omnibus and are accessible through GEO Series accession numbers (listed with the publications above).
Tracks for visualization in Xenbase G-Browse are available at the Xenbase FTP site.
Genome data publications
Akkers, R.C., S.J. van Heeringen, U.G. Jacobi, E.M. Janssen-Megens, K.J. Françoijs, H.G. Stunnenberg and G.J.C. Veenstra. 2009. A hierarchy of H3K4me3 and H3K27me3 acquisition in spatial gene regulation in Xenopus embryos. Developmental Cell, 17, 425-434.
![]() Dev.Cell Online
Dev.Cell Online
![]()
![]() PubMed Central
PubMed Central
![]() NCBI GSE14025
NCBI GSE14025
![]()
Van Heeringen, S.J., W. Akhtar, U.G. Jacobi, R.C. Akkers, Y. Suzuki, G.J.C. Veenstra. 2011. Nucleotide composition-linked divergence of vertebrate core promoter architecture. Genome Research, 21, 410-421.
![]() Genome Research Online
Genome Research Online
![]()
![]() NCBI GSE21482
NCBI GSE21482
![]()
Bogdanovic, O., S.W. Long, S. J. van Heeringen, A.B. Brinkman, J.L. Gómez-Skarmeta, H.G. Stunnenberg, P.L. Jones and G.J.C. Veenstra. 2011. Temporal uncoupling of the DNA methylome and transcriptional repression during embryogenesis. Genome Research, 21, 1313-1327 (doi: 10.1101/gr.114843.110).
![]() Genome Research Online
Genome Research Online
![]()
![]() NCBI GSE23913
NCBI GSE23913
![]()
Paranjpe SS, U.G. Jacobi, S.J. van Heeringen, G.J.C. Veenstra. 2013. A genome-wide survey of maternal and embryonic transcripts during Xenopus tropicalis development. BMC Genomics, 14,762 (doi:10.1186/1471-2164-14-762).
![]() BMC Genomics
BMC Genomics
![]() NCBI GSE43652
NCBI GSE43652
![]()
Heeringen, S.J. van, R.C. Akkers, I. van Kruijsbergen, M.A. Arif, L.L.P. Hanssen, N. Sharifi, G.J.C. Veenstra. 2014. Principles of nucleation of H3K27 methylation during embryonic development. Genome Research, 24, 401-410 (doi: 10.1101/gr.159608.113).
![]() Genome Research
Genome Research
![]() PDF
PDF
![]() NCBI GSE41161
NCBI GSE41161
![]()
